As the COVID-19 pandemic continues to evolve, it has become increasingly important to systematically adapt testing capacities to detect all variants of concern.
In a recently published article, Medical Lead for Microbiology and Virology at St. Paul’s Hospital, Dr. Marc Romney, and his team proposed a new testing strategy that can rapidly detect circulating variants, and which enabled them to uncover a large swath of cases of a variant B.1.1.28/P.1, the one originating from Japan/Brazil, previously undocumented in British Columbia. What’s more, this new method is quicker and cheaper than the current standard, does not require samples carrying high concentrations of virus, and can be scaled-up to test all SARS-CoV-2 positive samples.
The early and rapid detection of SARS-CoV-2 variants of concern is of utmost public health importance to prevent and control widespread community transmission of troublesome variants. Not only have these variants evolved to become more transmissible and less susceptible to neutralizing antibodies, but their multiple newly-acquired mutations could also make them harder to capture using certain conventional testing methods.
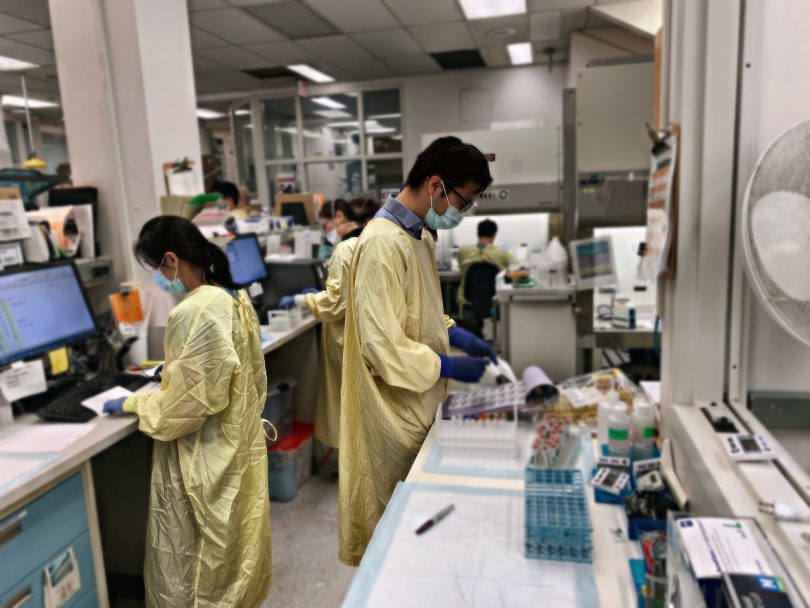
Samples analyzed at St. Paul’s virology lab
The authors of this study analyzed nearly 32,000 nasal swabs and saliva/mouth rinse samples collected between January 25 and March 1, 2021 in BC. Approximately eight per cent came back positive for SARS-CoV-2. The majority of positive samples underwent further testing to identify and track the spread of variants of concern, specifically lineages B.1.1.7, B.1.351 and B.1.1.28/P.1, first identified in the United Kingdom, South Africa and Japan/Brazil, respectively. This was accomplished using an assay that scans the genomic sequence of the virus for a mutation called N501Y that is common to all current variants of concern but lacking in the standard strain of SARS-CoV-2. Samples harbouring this worrisome mutation were then further analyzed to distinguish amongst variants of concern. They were examined for additional mutations that are known to be specific to each variant.
Ultimately, 77 variants of concern were identified, 57 of which matched to the variant originating from the UK, seven to the one originating from South Africa, and 13 to the one originating from Japan/Brazil. The latter finding was rather unanticipated seeing as at the time of the study, there were very few cases of this troublesome variant circulating in Canada and the United States, and none previously detected in BC.

“Using this technology, we rapidly identified a cluster of B.1.1.28/P.1 cases which otherwise would have likely gone undetected,” says Dr. Romney. “Since then, we have identified more B.1.1.28/P.1 cases in BC than have been identified in the entire United States, and more than any other country, except Brazil and Italy.”
Testing strategy is faster and cheaper
Importantly, variants were identified within 24-hours of analysis, which is a more rapid turnaround time than one of the current variant surveillance strategies called whole genome sequencing. In Canada, whole genome sequencing is performed on a random subgroup of samples positive for SARS-CoV-2, meaning certain cases of variants could go unnoticed. And not only is this method time-consuming, but it is also quite expensive.
On the other hand, a PCR-based algorithm, such as the one presented by Dr. Romney and colleagues, is cheaper and can easily be scaled-up to screen all positive samples in a diagnostic laboratory, thereby giving a head start to the appropriate public health follow-up (contact tracing, outbreak declaration, etc.). Additionally, PCR-based testing strategies are adaptable and can be updated on occasion to include targets of newly emerging strains of the virus. Finally, this approach also does not require the presence of large amounts of virus to be present in the clinical specimen.
The authors suggest that their testing strategy would be a great addition to the current method of ongoing variant strain surveillance performed in SARS-CoV-2 diagnostic laboratories today.
This story originally appeared on the COVID-19 Immunity Task Force (CITF) website and was adapted with permission. CITF provided partial funding for this study. Read the original story here.